Efficient target cleavage by Type V Cas12a effectors programmed with split CRISPR RNA
Nucleic Acids Res. 2021 Dec 24. pii: gkab1227. doi: 10.1093/nar/gkab1227. Online ahead of print.
Regina Shebanova 1, Natalia Nikitchina 1 2, Nikita Shebanov 1, Vladimir Mekler 3, Konstantin Kuznedelov 3, Egor Ulashchik 4, Ruslan Vasilev 5 6, Olga Sharko 4, Vadim Shmanai 4, Ivan Tarassov 2, Konstantin Severinov 1 3 7, Nina Entelis 2, Ilya Mazunin 1
Abstract
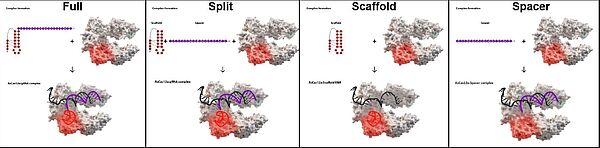
CRISPR RNAs (crRNAs) that direct target DNA cleavage by Type V Cas12a nucleases consist of constant repeat-derived 5'-scaffold moiety and variable 3'-spacer moieties. Here, we demonstrate that removal of most of the 20-nucleotide scaffold has only a slight effect on in vitro target DNA cleavage by a Cas12a ortholog from Acidaminococcus sp. (AsCas12a). In fact, residual cleavage was observed even in the presence of a 20-nucleotide crRNA spacer moiety only. crRNAs split into separate scaffold and spacer RNAs catalyzed highly specific and efficient cleavage of target DNA by AsCas12a in vitro and in lysates of human cells. In addition to dsDNA target cleavage, AsCas12a programmed with split crRNAs also catalyzed specific ssDNA target cleavage and non-specific ssDNA degradation (collateral activity). V-A effector nucleases from Francisella novicida (FnCas12a) and Lachnospiraceae bacterium (LbCas12a) were also functional with split crRNAs. Thus, the ability of V-A effectors to use split crRNAs appears to be a general property. Though higher concentrations of split crRNA components are needed to achieve efficient target cleavage, split crRNAs open new lines of inquiry into the mechanisms of target recognition and cleavage and may stimulate further development of single-tube multiplex and/or parallel diagnostic tests based on Cas12a nucleases.